CRC 1430
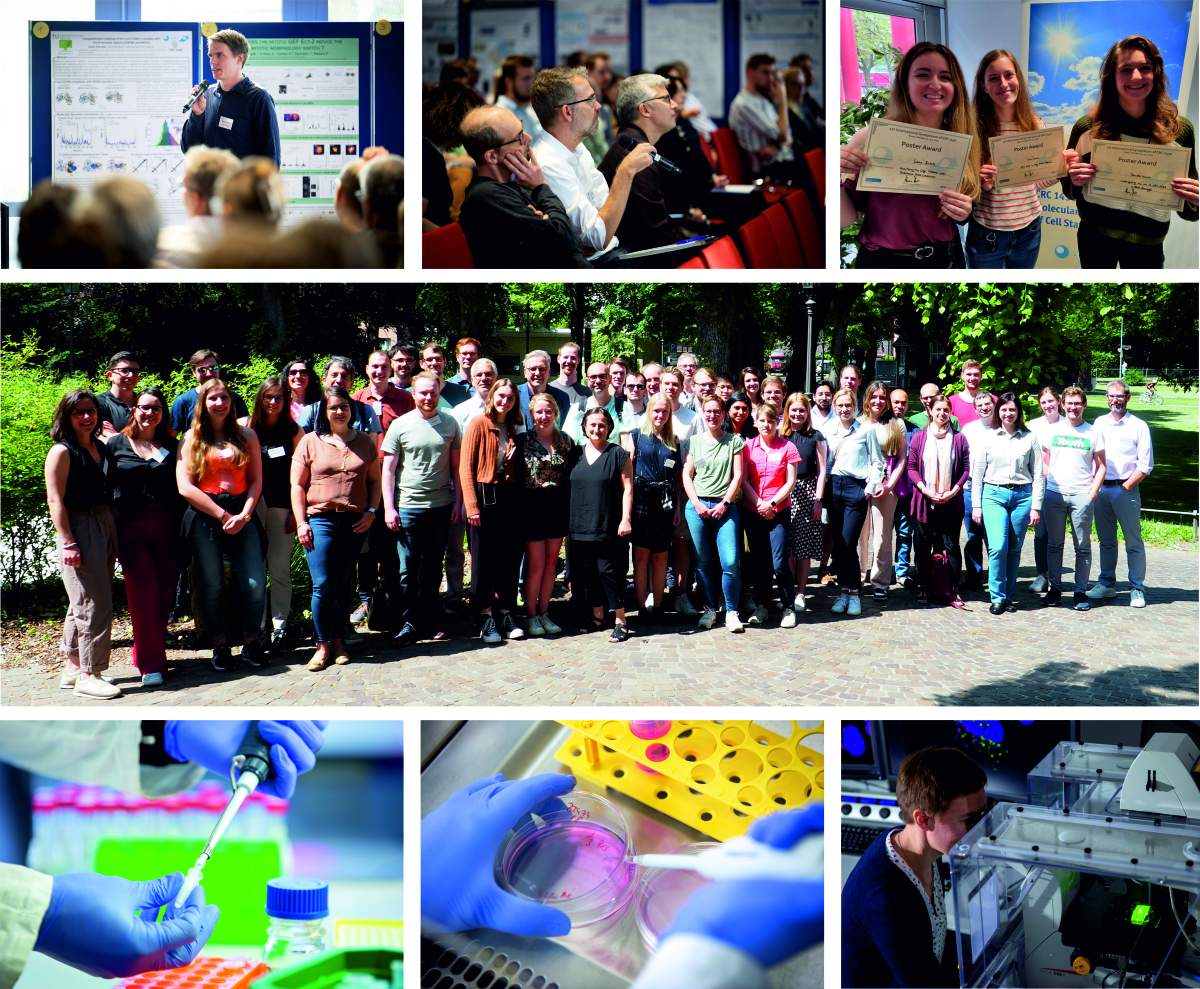
Welcome to the Collaborative Research Centre "Molecular Mechanisms of Cell State Transitions" (CRC 1430)
The DFG-funded CRC 1430 "Molecular Mechanisms of Cell State Transitions" explores fundamental molecular mechanisms that underlie the regulation of cell proliferation. Cell proliferation needs to be tightly controlled to ensure organismal development and tissue regeneration, while preventing neoplastic disorders. A key hallmark of this control is the establishment of distinct, biochemically or epigenetically defined cell states and the regulated transitions between these states.
These transitions govern cell cycle progression and underlie cancer cell plasticity and cancer therapy resistance. The research focus is on understanding the switch-like molecular trigger mechanisms of state transitions and develop means to modulate them, ultimately to identify novel therapeutic strategies. Specifically, to overcome current limitations, the CRC 1430 will develop and apply direct methodologies such as advanced biochemical reconstitution and novel approaches of acute chemical or optical perturbation to decipher how the key triggers sense, integrate and transmit signals to regulatory circuits that define cell states.

Upcoming ZMB/CRC 1430 Guest Lectures
| 13.10.2026 | Helle Ulrich Ubiquitin, SUMO and genome maintenance, IMB Mainz |
A list of previous speakers can be found here:

Upcoming CRC1430 Workshops






Partner