CRC 1093 - Overview
General concepts I
The two following schematics illustrate which supramolecular approaches the chemical groups will exploit (blue), in order to manipulate proteins investigated in the biological groups (yellow). This picture focusses on novel methods of protein recognition with supramolecular synthetic binders developed within projects A1-A3, A10 and A11 the addressed protein targets B1-B7. Theoretical treatment and advanced analysis in A7-A9.
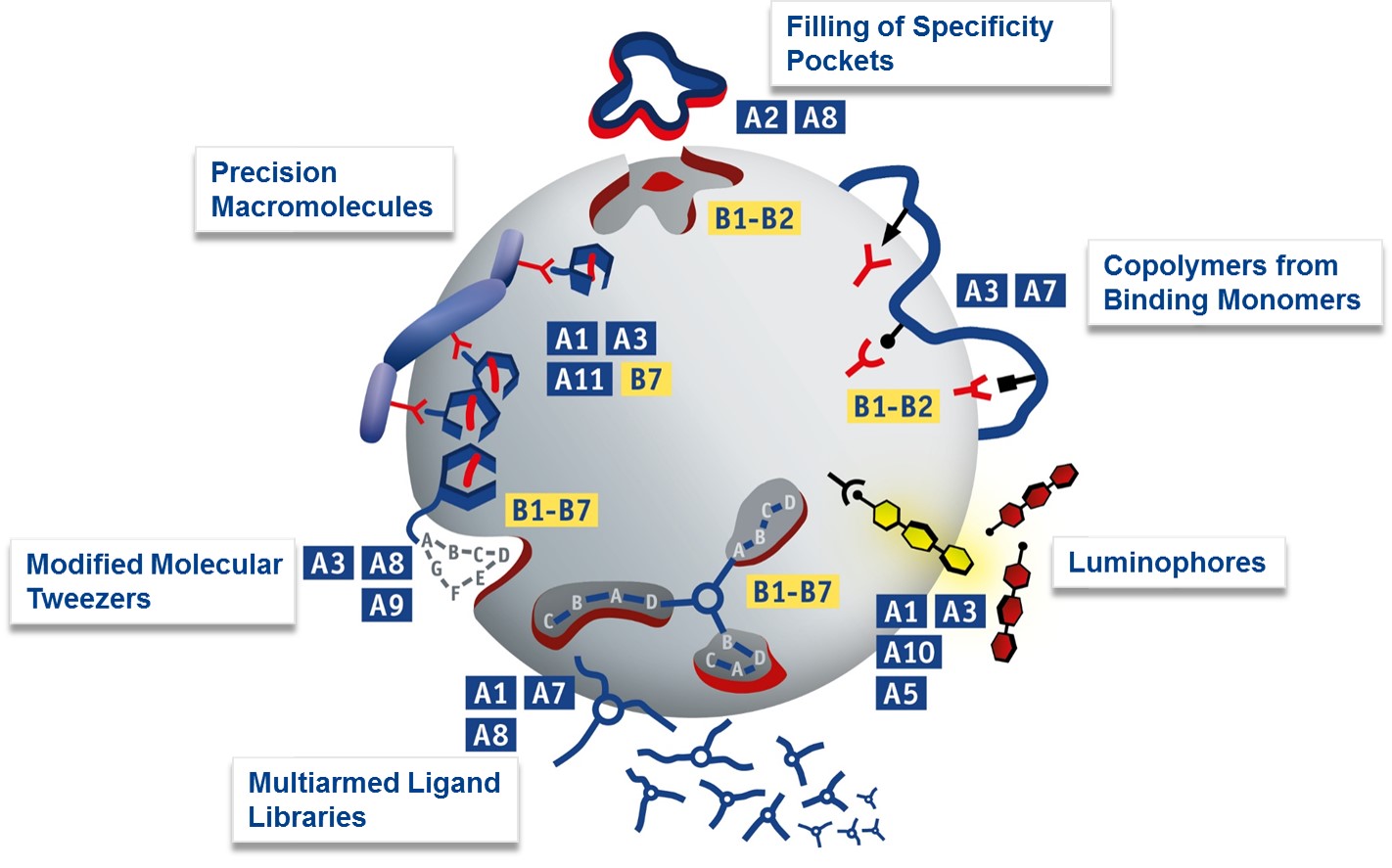
A2/A8: Protein-selective natural product obtained by filling of specificity pockets; A3/A7: Affinity polymers from amino acid-selective comonomers; A10/A1/A3: Light-up Luminophors; A1/A7/A8: Triarmed ligand library (DCC method); A3/A8/A9: Molecular lysine tweezers equipped with epitope recognition motif. A11/A1/A3: Precision oligomers for multivalent presentation of supramolecular ligands. A7: Prediction of maximal heteroavidity by Pareto optimization; A8: Molecular simulation of large protein/ligand complexes by MD-QM/MM combination. A9: Site-specific quantification of protein ligand interactions with Raman techniques.
B1-B2 (top and right): Serine proteases addressed by tailored active site or protein surface binders. B1-B7 (bottom): Proteins with extended recognition domains for combinatorial screening with focussed libraries; B1-B7 (bottom left): Proteins with well accessible basic or acidic residues in the vicinity of critical binding epitopes for peptide or protein partners.
General concepts II
Protein interactions examined in B1-B7 which are targeted with supramolecular ligands from projects A1-A11 supported by theoretical treatment and advanced analysis in A7-A9.
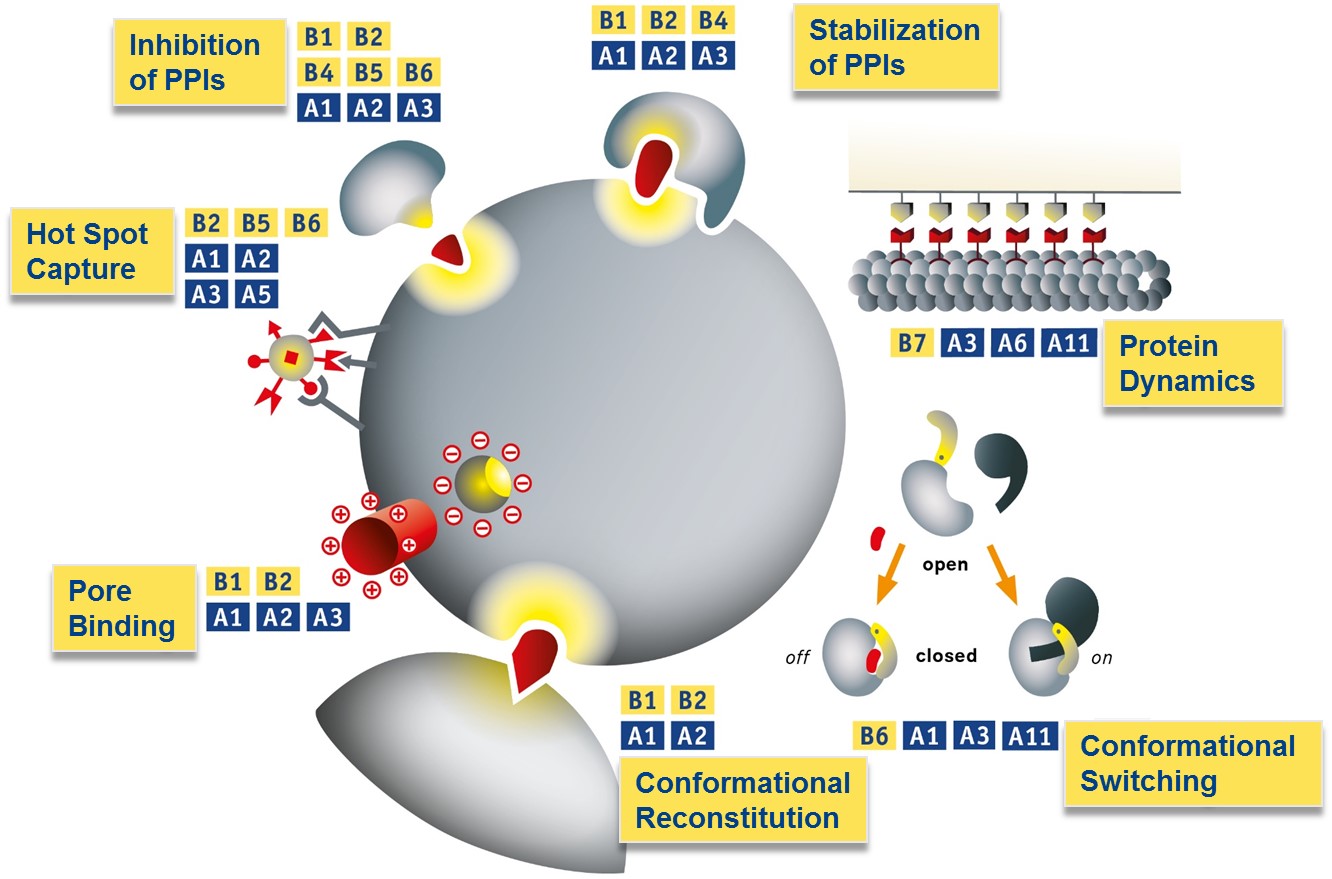
B1-B4/A1-A3 (top): Supramolecular Stabilization of PPIs; B7/A3/A6/A11 (right): Microtubule recognition with multivalent ligands using DNA-origami architectures; B6/A1/A3/A11 (bottom right): Switching between protein conformations; B1/B2/A1/A2 (bottom): Restoration of pathogenic protein conformations; B1/B2/A1-A3 (center): Molecular Corks for Protein Pores; B2-B6/A1-A3/A5 (left): Hot spot targeting by ultrasmall functional nanoparticles; B1-B6/A1-A3(top left): Supramolecular Inhibition of PPIs.
Area Z: Central Service
Z2: X-ray crystallography for all groups of the CRC. Cocrystallization and soak-in experiments. Mechanistic and structural elucidation of protein ligand complexes.
Z3: NMR spectroscopy for all groups of the CRC. Ligand optimization (STD), epitope identification (chemical shift perturbation) and structural characterization of protein ligand complexes (TROSY).